PartSeg
A GUI and a library for segmentation algorithms.
PartSeg
PartSeg is a GUI and a library for segmentation algorithms. PartSeg also provide napari plugins for IO and labels measurement.
This application is designed to help biologist with segmentation based on threshold and connected components.
Tutorials
- Tutorial: Chromosome 1 (as gui) link
- Data for chromosome 1 tutorial link
- Tutorial: Different neuron types (as library) link
Installation
From binaries:
With pip:
- From pypi:
pip install PartSeg[all] - From repository:
pip install git+https://github.com/4DNucleome/PartSeg.git
- From pypi:
With conda:
conda install -c conda-forge partsegmamba install -c conda-forge partseg- As mamba is faster than conda
With napari:
If you do not know how to setup python environment on your system you may use napari to run PartSeg. It is a GUI for scientific image analysis. PartSeg is also a plugin for napari so could be installed from plugin dialog. To install napari bundle please download it napari bundle and follow installation instructions.
Installation troubleshooting information could be found in wiki: wiki. If this information does not solve problem you can open issue.
Qt 6 support
PartSeg development branch support (and stable since 0.15.0) has experimental Qt6 support. Test are passing but not whole GUI code is covered by tests. Inf you Find any problem please report it.
Running
If you downloaded binaries, run the PartSeg (or PartSeg.exe for Windows) file inside the PartSeg folder
If you installed from repository or from pip, you can run it with PartSeg command or python -m PartSeg.
First option does not work on Windows.
PartSeg export few commandline options:
--no_report- disable error reporting--no_dialog- disable error reporting and error dialog. Use only when running from terminal.roi- skip launcher and start ROI analysis guimask- skip launcher and start ROI mask gui
napari plugin
PartSeg provides napari plugins for io to allow reading projects format in napari viewer.
Save Format
Saved projects are tar files compressed with gzip or bz2.
Metadata is saved in data.json file (in json format). Images/masks are saved as *.npy (numpy array format).
Interface
Launcher. Choose the program that you will launch:
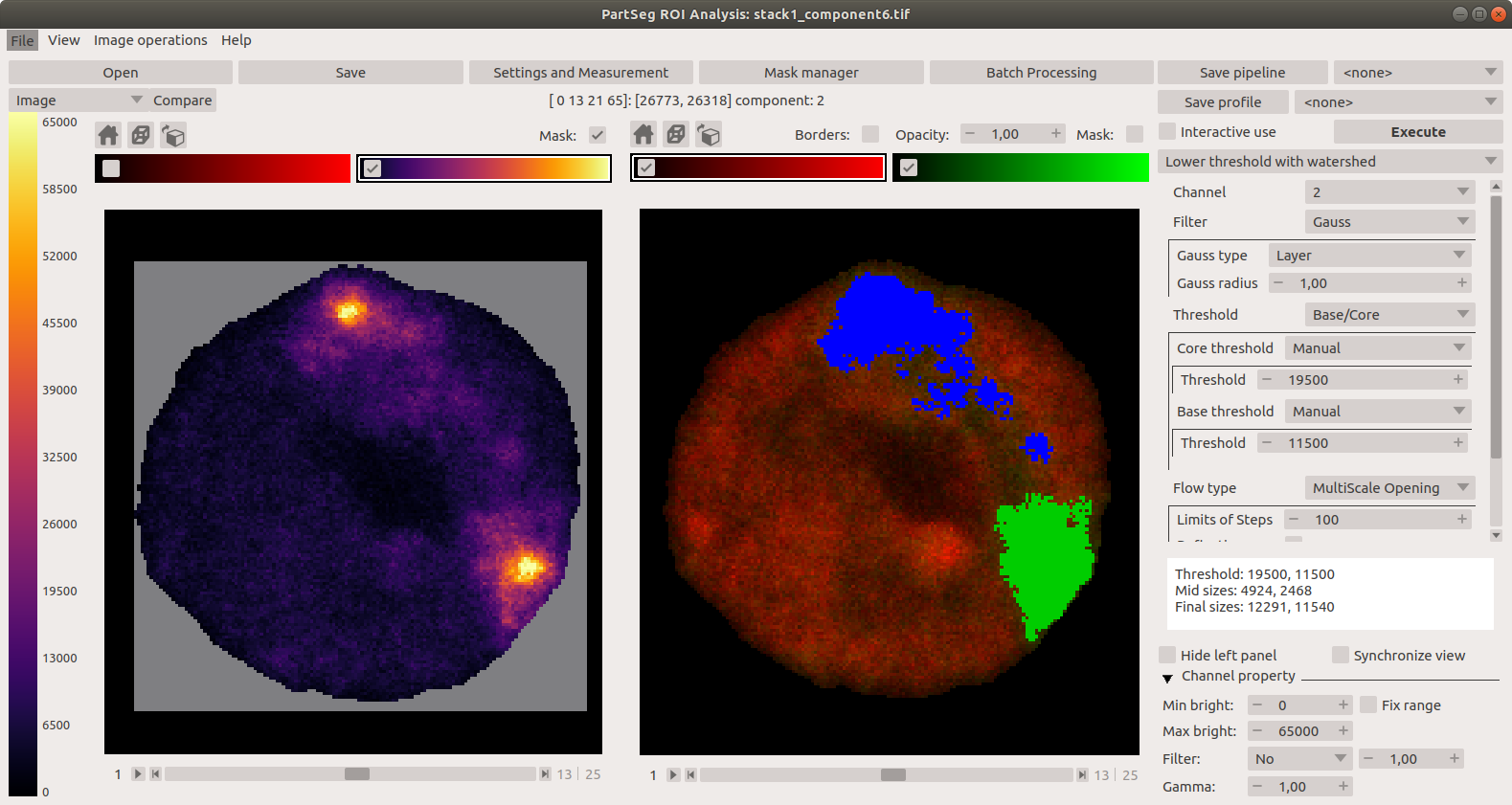
Main window of Segmentation Analysis:

Main window of Segmentation Analysis with view on measurement result:
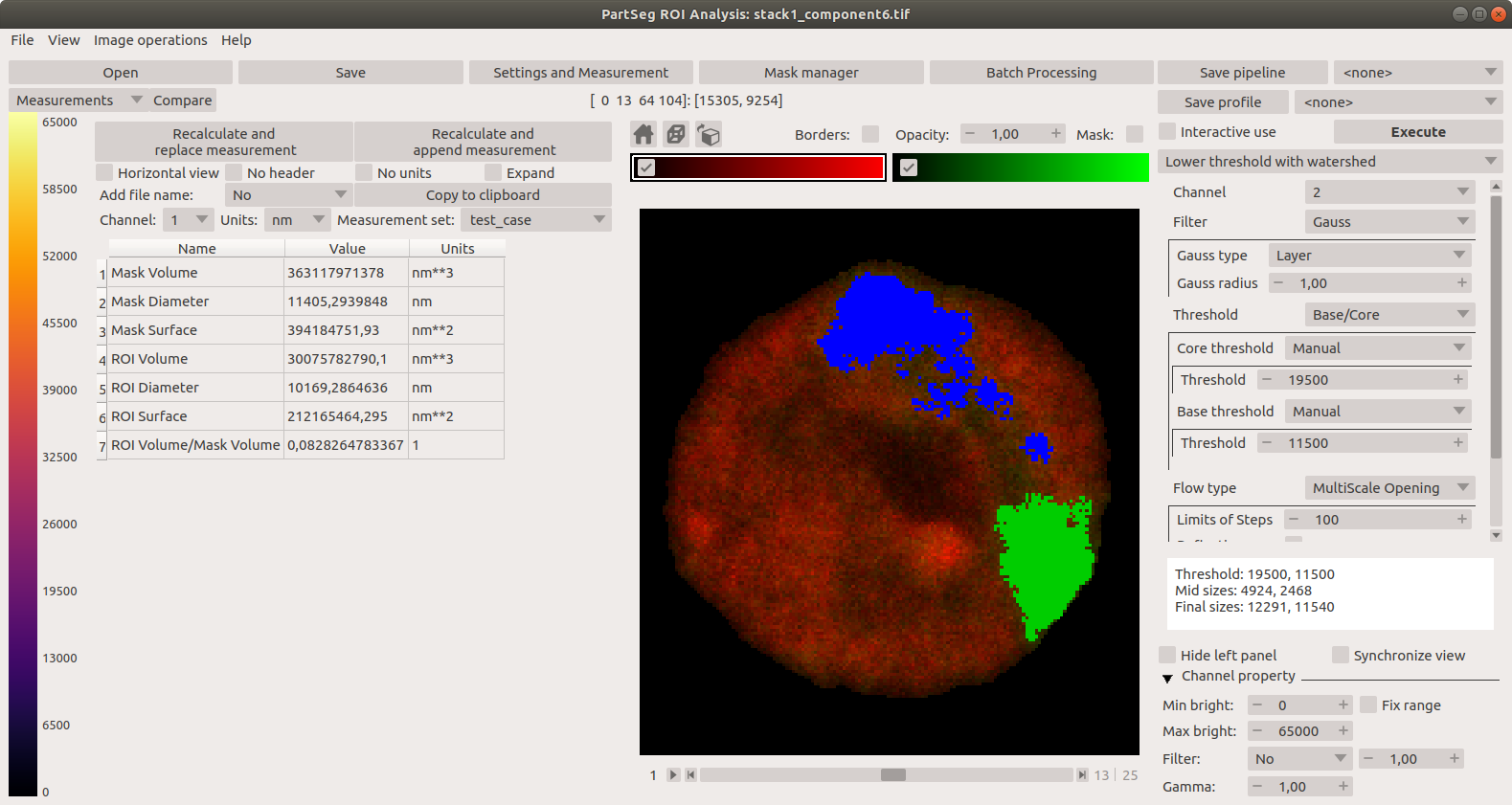
Window for creating a set of measurements:
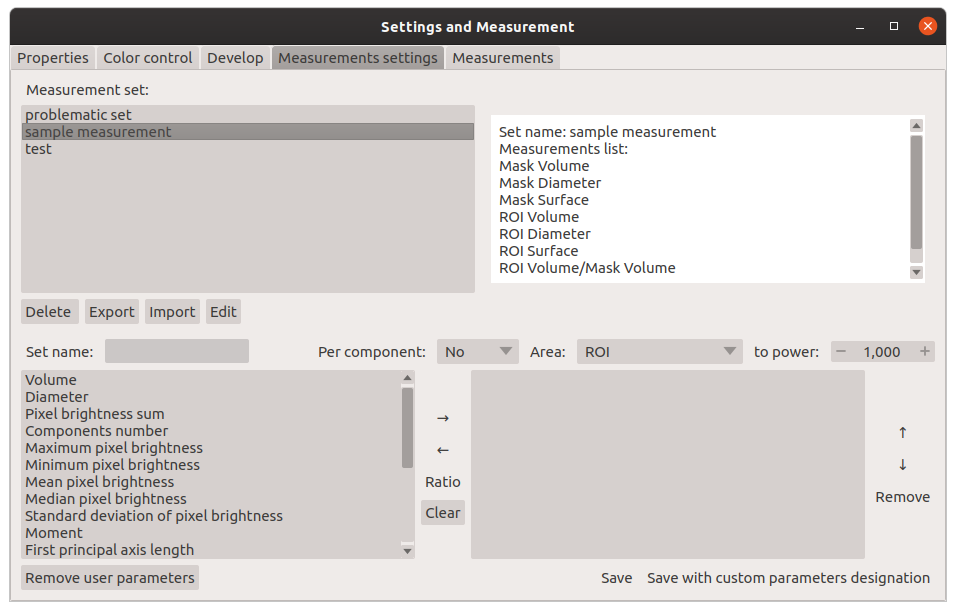
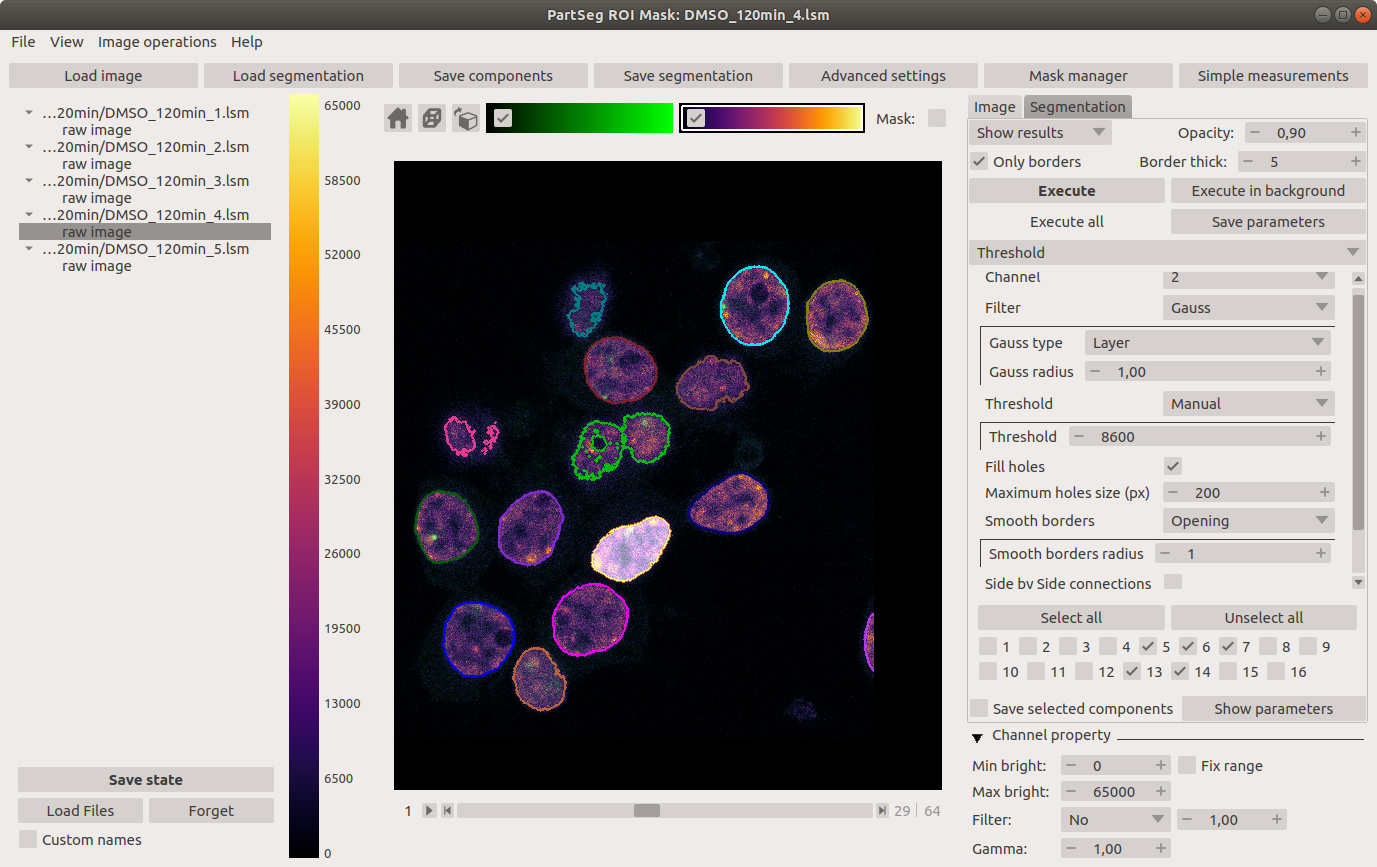
Main window of Mask Segmentation:
Laboratory
Laboratory of Functional and Structural Genomics http://4dnucleome.cent.uw.edu.pl/
Cite as
Bokota, G., Sroka, J., Basu, S. et al. PartSeg: a tool for quantitative feature extraction from 3D microscopy images for dummies. BMC Bioinformatics 22, 72 (2021). https://doi.org/10.1186/s12859-021-03984-1






