Spatial Foundation Models > scGPT-spatial
Extension of scGPT for spatial transcriptomics with continual pretraining and a mixture-of-experts decoder for spatial gene expression analysis.
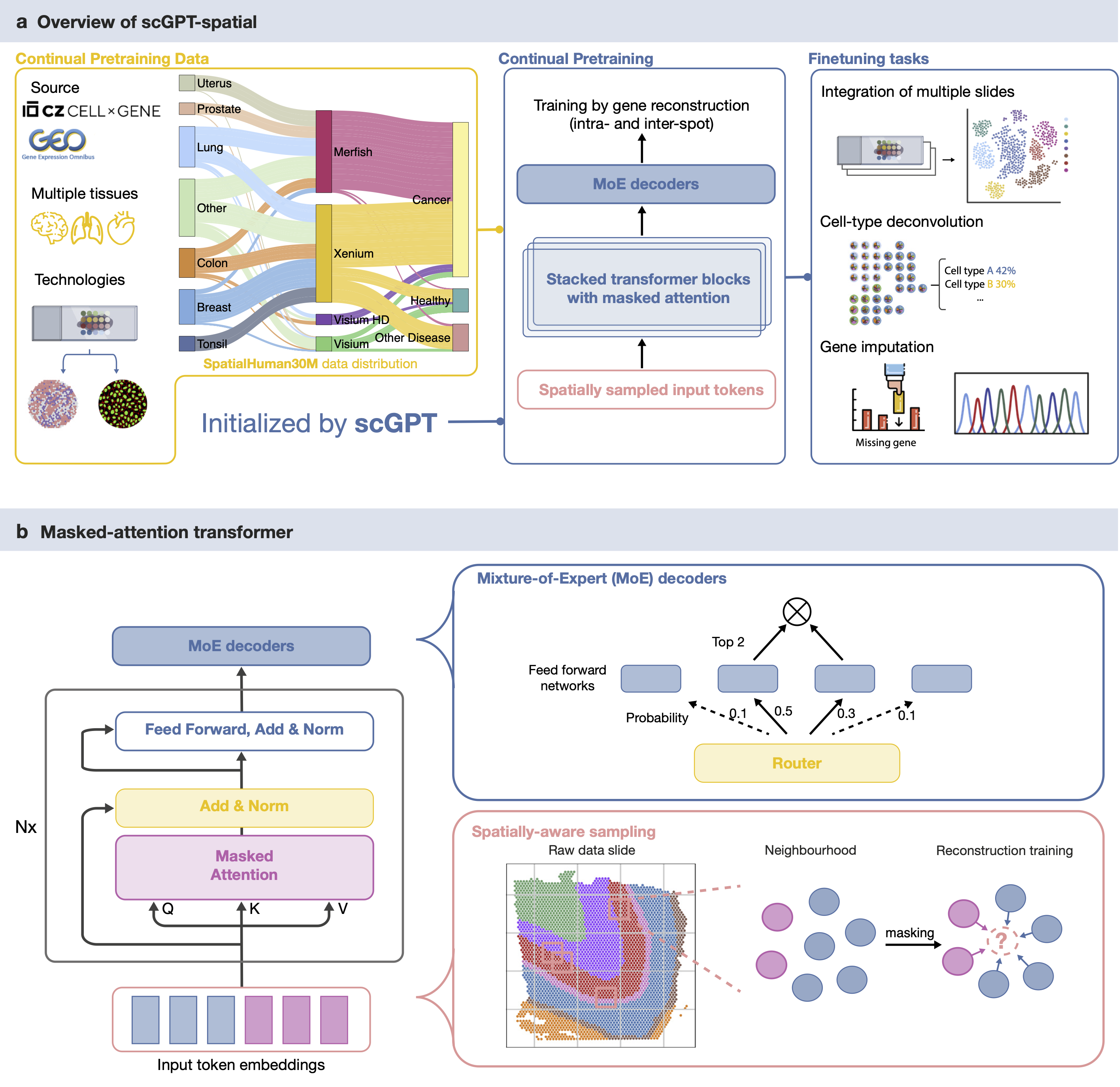
scGPT-spatial: Continual Pretraining of Single-Cell Foundation Model for Spatial Transcriptomics
🟩 TL,DR Highlights 🟩
✨ Spatial-omic foundation model ✨ ✨ Continual pretraining of scGPT on 30 million cells/spots ✨
✨ Novel MoE (Mixture of Experts) decoders ✨ ✨ Spatially-aware sampling ✨ ✨ Neighborhood-based reconstruction objective ✨
✨ Curation of SpatialHuman30M corpus ✨ ✨ Visium, Visium HD, Xenium, MERFISH ✨
✨ Multi-modal and multi-slide integration ✨ ✨ Cell-type deconvolution ✨ ✨ Missing gene imputation ✨
🟧 Model Weights 🟧
scGPT-spatial V1 weights on figshare.
🟫 SpatialHuman30M 🟫
Pretraining dataset names, slide metadata, and access links are summarized in data source table. Processed data will be available upon publication given permission under license of the original data source.
🟦 Setup and Tutorials 🟦
To start, clone the current repo:
git clone https://github.com/bowang-lab/scGPT-spatial
Special acknowledgement to the scGPT codebase - for environment setup please follow instructions there.
Check out our zero-shot inference tutorial on github! More code coming soon.
🟪 Preprint and Citation 🟪
Check out our preprint! https://www.biorxiv.org/content/10.1101/2025.02.05.636714v1
@article{wang2025scgpt,
title={scGPT-spatial: Continual Pretraining of Single-Cell Foundation Model for Spatial Transcriptomics},
author={Wang, Chloe Xueqi and Cui, Haotian and Zhang, Andrew Hanzhuo and Xie, Ronald and Goodarzi, Hani and Wang, Bo},
journal={bioRxiv},
pages={2025--02},
year={2025},
publisher={Cold Spring Harbor Laboratory}
}