Drug Response Prediction > cycleCDR
Interpretable cycle-consistency framework for modeling cellular responses to drug perturbations.
Interpretable modeling of cellular responses to drug perturbations using cycle consistence learning
Introduction
cycleCDR is interpretable modeling of cellular responses to drug perturbations using cycle consistence learning.
Model architecture
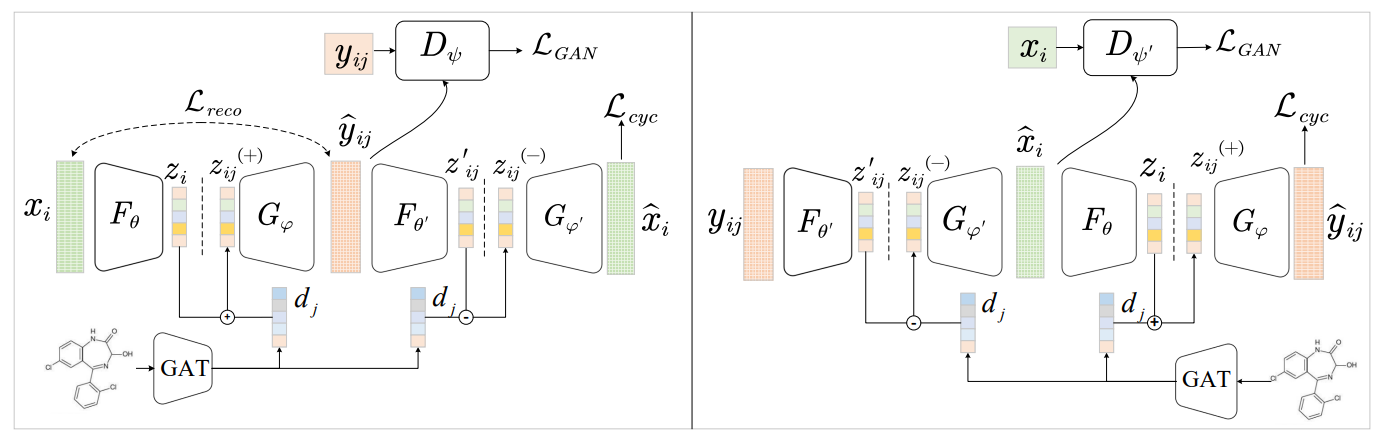
Requirements
The deep learning models were trained on 2*NVIDIA GeForce RTX 4090 on linux.
- Python 3.11
- PyTorch 2.0
- Pandas 2.1.1
- Numpy 1.25.2
- Scikit-learn 1.3.1
Usage
To setup the environment, install conda and run (Must run on servers with multiple GPUs):
conda create --name <your_env_name> --file requirements.txt
To train this model and obtain results, you need to download the dataset (Example: L1000), place it in the datasets folder, and then run:
torchrun --nproc_per_node=2 --master-port=29501 cycleCDR/linc_main.py
If you want to train other datasets, you need to modify the cycleCDR/linc_main.py section
Directory structure
cycleCDR: contains the code for the model, the dataset, the evaluation, and the training loop.preprocessing: Scripts for processing the data.configs: Configuration file for hyperparameters.datasets: Directory where data is stored.results: This directory needs to be created by oneself. It is used to store the results of the experiment.modules: Model storage directory.plot: Training loss curve.plot_data: Data of the training process.